Getting started on the web
The GAIn Web Annotation page (https://gain.iossifovlab.com/) provides a simple and interactive interface for annotating genetic variants, positions, or regions, collectively referred to as annotatables, on the web. The page is organized into two main areas. On the left, users can create, edit, and manage annotation pipelines, either by selecting a saved pipeline or by building a new one. This pipeline editor defines which resources and annotators are applied during annotation. On the right, users can enter an annotatable, submit it for processing, and view the resulting annotations in the same area. Together, these two panels allow users to configure an annotation workflow and immediately apply it to annotatables of interest.
Select annotation pipeline
To begin, open the Select pipeline menu, scroll through the available options, and choose
pipeline/hg38_Clinical_annotation. This pipeline is designed for annotatables provided in the hg38
assembly and annotates them with a broad set of clinically relevant attributes. In addition to effect prediction,
it incorporates information from resources such as dbSNP, gnomAD,
ClinVar, CADD, AlphaMissense, MPC, and gene-level constraint scores. After selecting it, users can review
the full annotation pipeline definition in the box on the left.
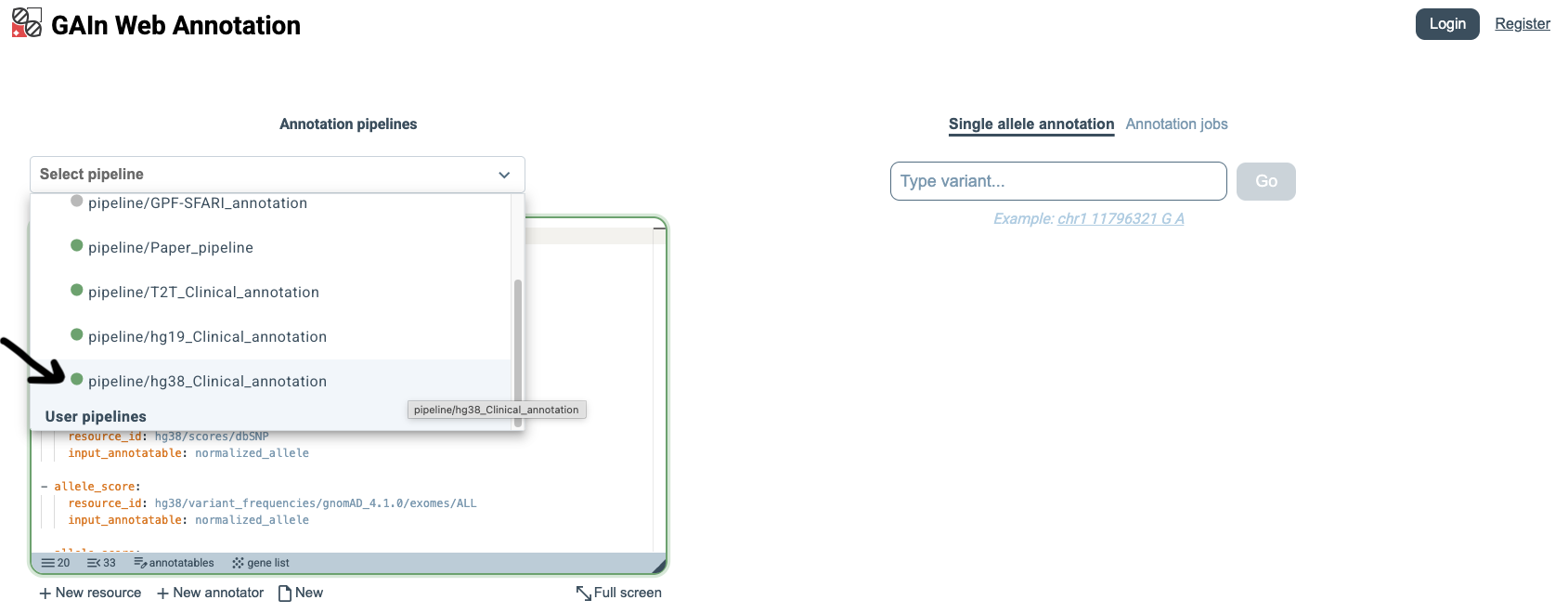
Single annotatable
The GAIn web interface has two annotation modes: Single annotatable, for interactively annotating one variant,
position, or region at a time, and Annotation jobs, for annotating files containing multiple annotatables.
We begin with the Single annotatable mode.
Next, either enter your own annotatable in hg38 coordinates, or click the i button left of the entry field and
choose an example annotatable.
Example annotatables cover a wide range of syntax options for entering genomic positions, regions, or variants.
This automatically enters the example annotatable, runs the selected pipeline on it, and displays the resulting
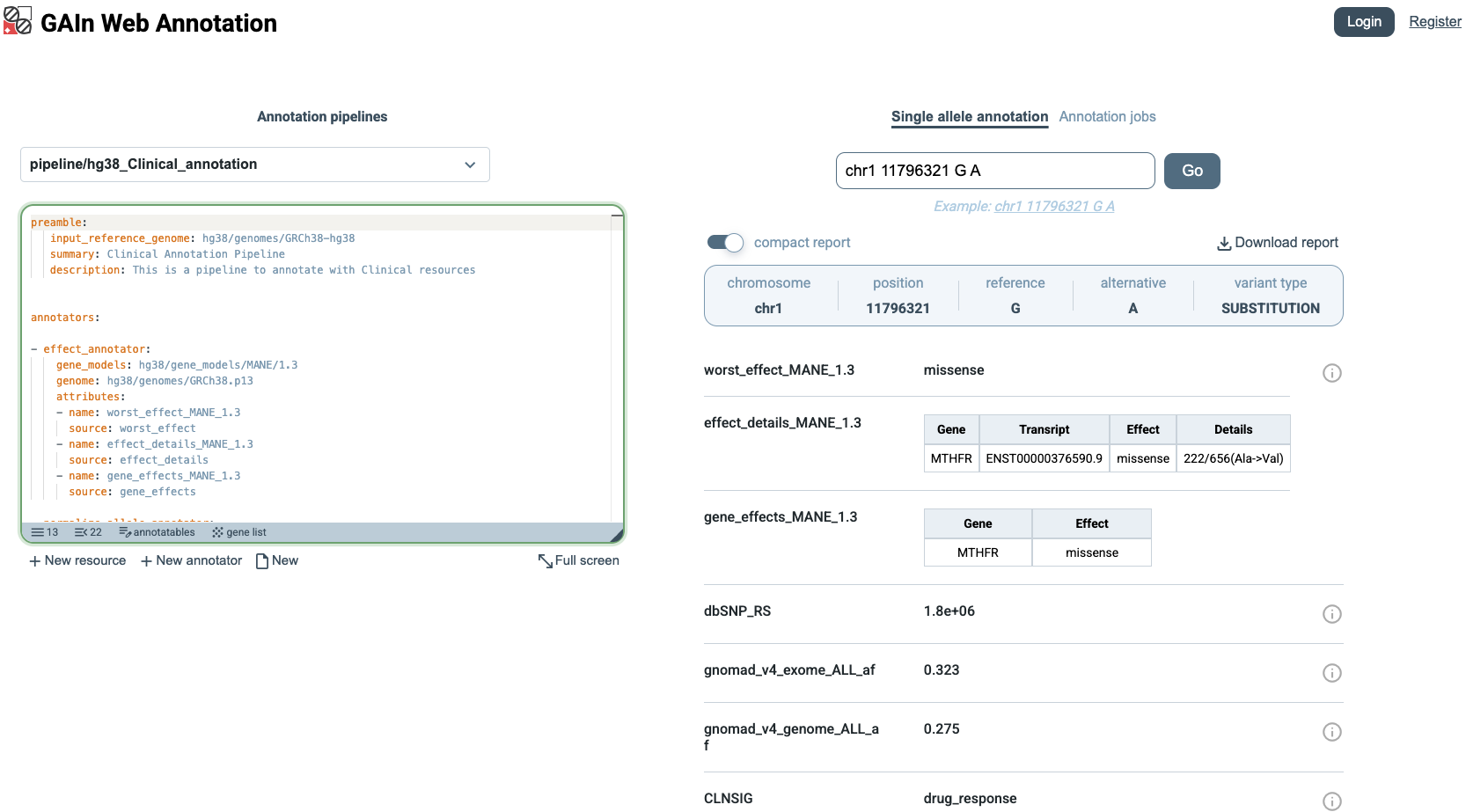
annotations on the right side of the page. In this example, we choose a variant annotatable,
which is defined by chromosome, position, reference allele, and alternative allele.
The annotation results can be viewed in two formats. In the compact report view, only the annotation results are shown, making it easier to scan the output. This view is useful when a concise summary is needed without additional details.
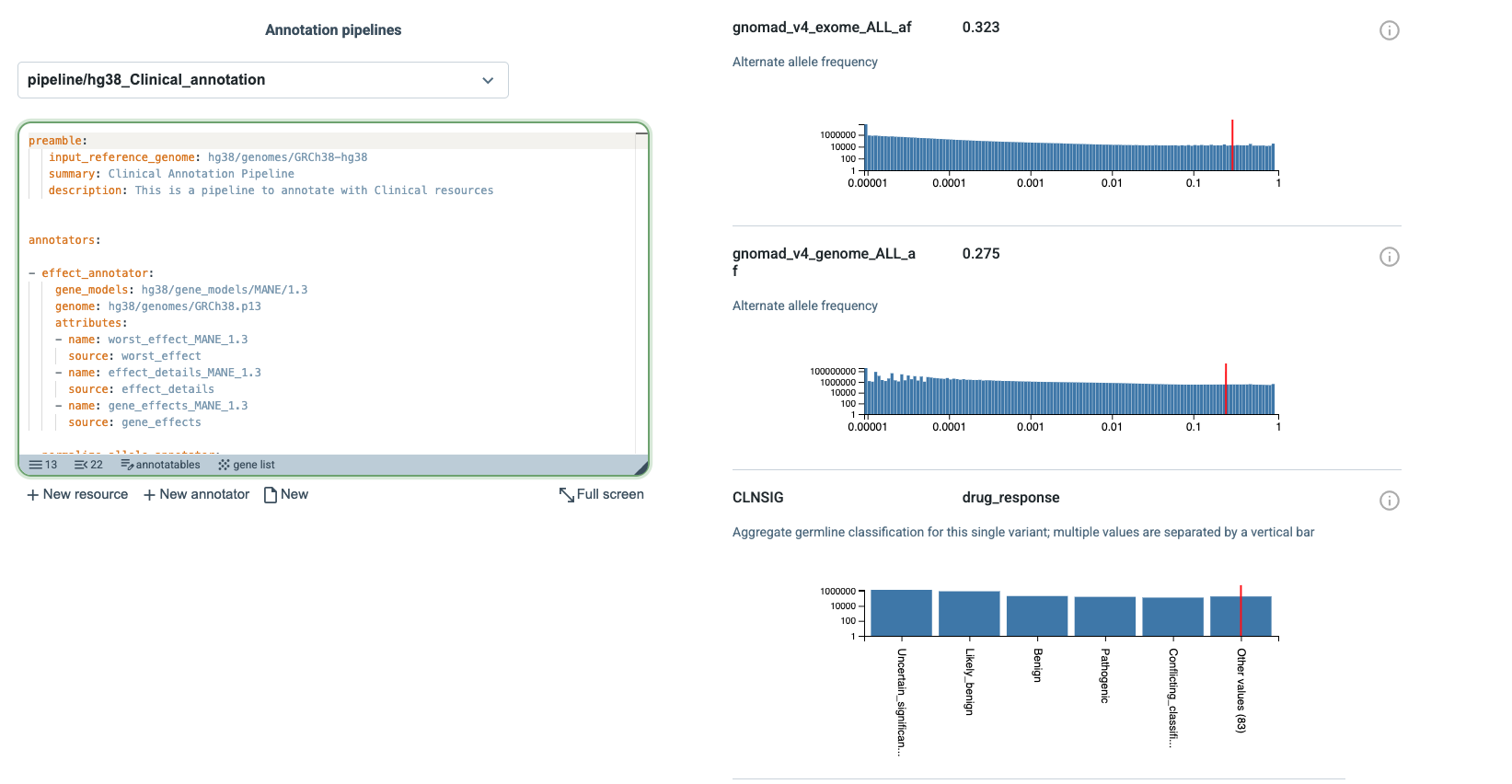
When compact report is turned off, the page shows a more detailed view of the results. In this view, annotations that include score distributions are accompanied by plots showing how the scores are distributed across the resource and where the queried annotatable falls within that distribution. In the example shown here, three such distributions are displayed. The red line marks the score of the queried annotatable, making it easy to see its position relative to the overall distribution.
Annotation jobs
The second annotation mode is Annotation jobs, which allows multiple annotatables to be annotated at once. In this mode, users upload a file in tabular or VCF format containing annotatables (variants, positions, or regions) with the required coordinate information. Click Annotation jobs on the right side of the page.
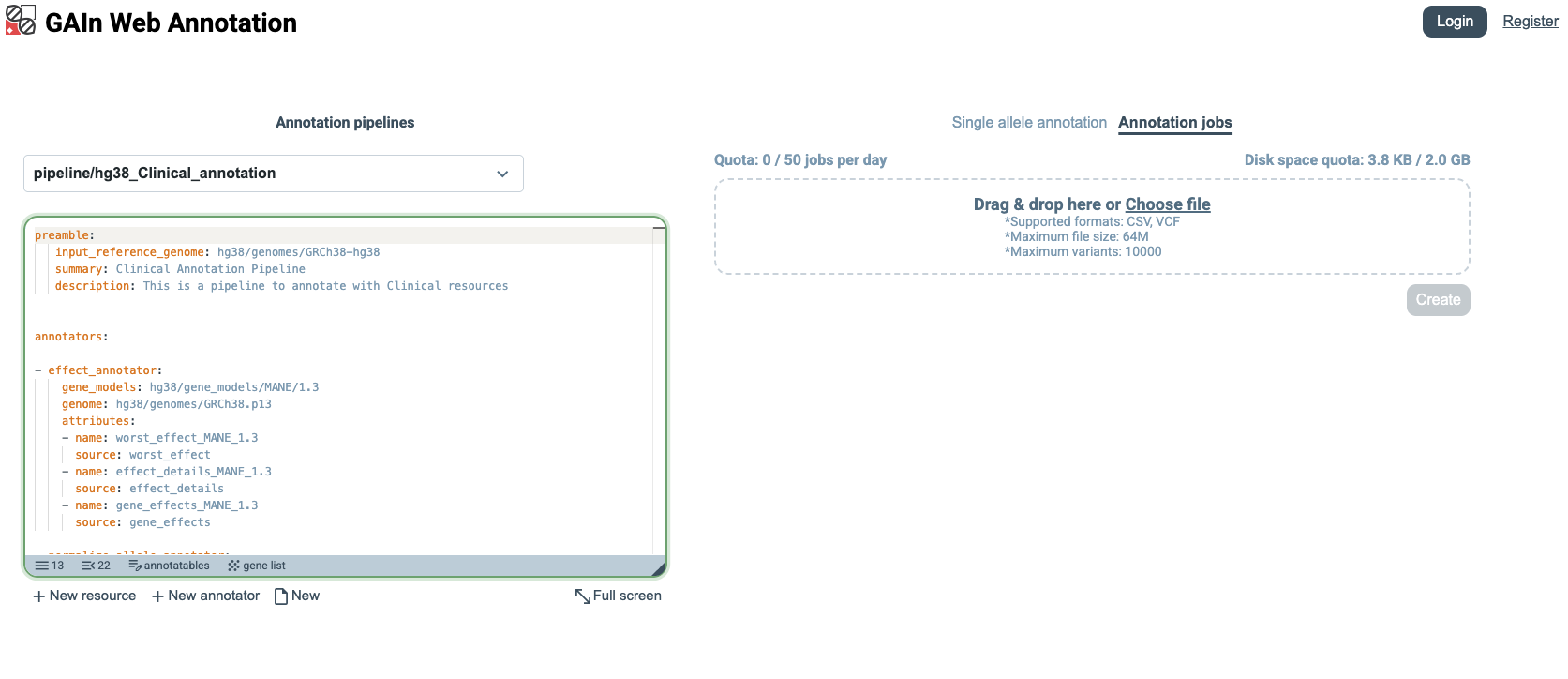
The example below uses variant annotatables.
Prepare a tab-delimited file named variants.txt with the following content:
chrom |
pos |
ref |
alt |
|---|---|---|---|
chr11 |
5227002 |
T |
A |
chr12 |
102840493 |
G |
A |
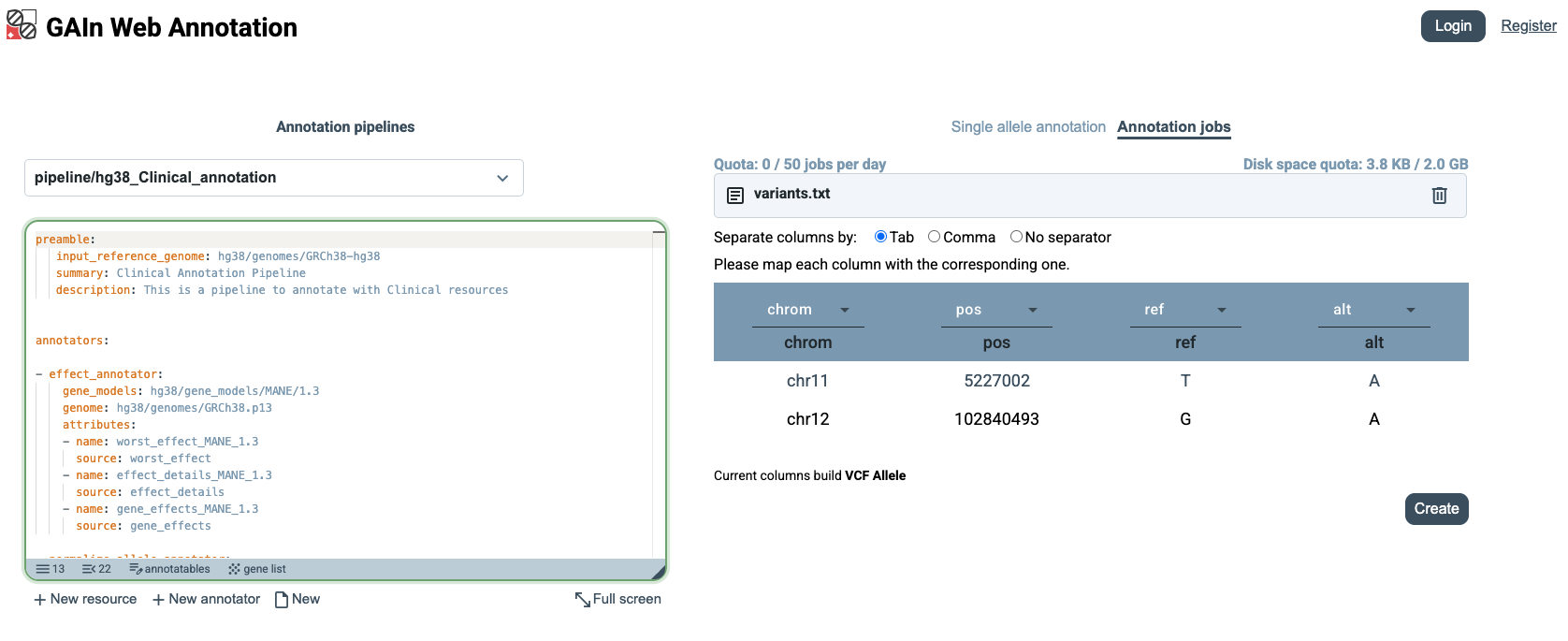
After creating this file, drag and drop it into the upload area, or click Choose file and select variants.txt for annotation.
When the file is uploaded, GAIn automatically recognizes the chrom, pos, ref, and alt columns because the file uses those exact column names. If the input file uses different column labels, users can manually specify which columns correspond to chrom, pos, ref, and alt before creating the annotation job.
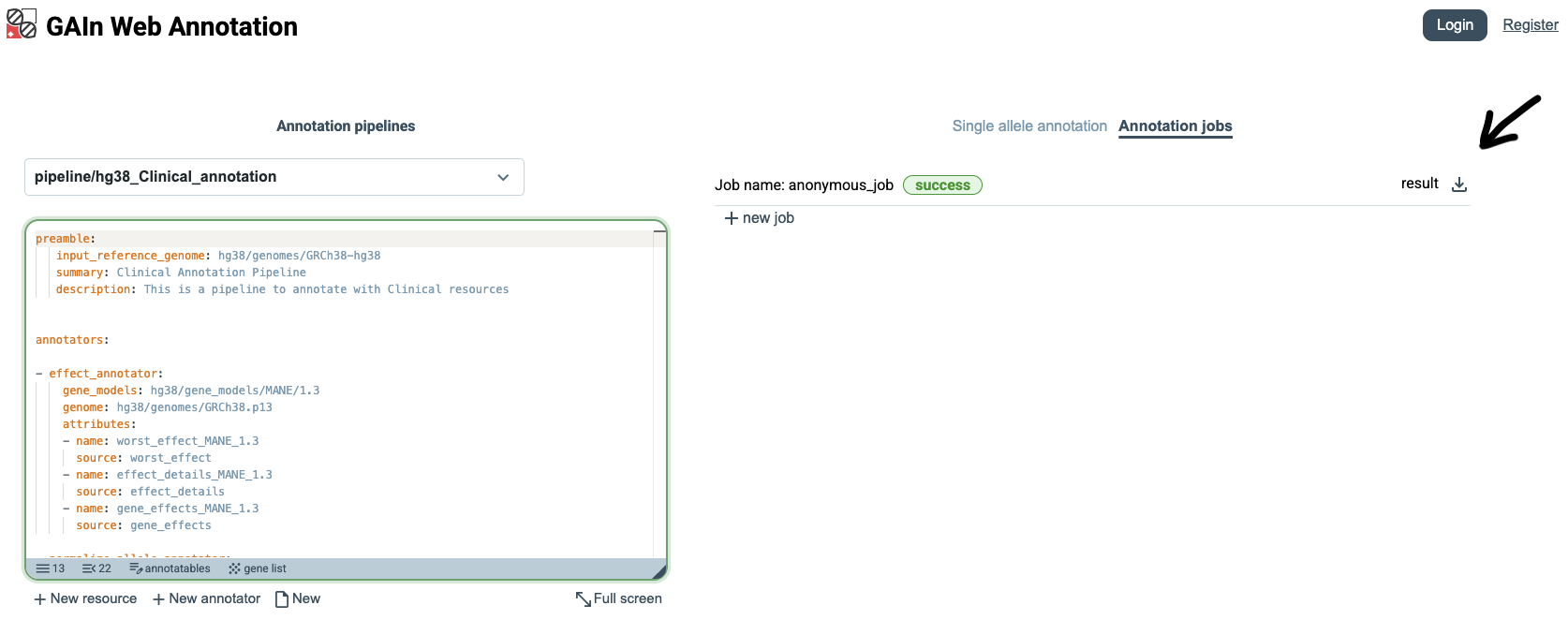
Once the annotation job has finished, GAIn shows its status as success.
Users can then click the Download button next to the result to download a file containing
the annotation attributes produced for all annotatables in the input file.
The downloaded file includes the original variant columns together with the annotations generated by
the selected pipeline,
such as effect predictions, population frequencies, clinical classifications, pathogenicity scores,
and gene-level scores. The downloaded output for variants.txt is shown below.
chrom |
pos |
ref |
alt |
worst_effect_MANE_1.3 |
effect_details_MANE_1.3 |
gene_effects_MANE_1.3 |
dbSNP_RS |
gnomad_v4_exome_ALL_af |
gnomad_v4_genome_ALL_af |
CLNSIG |
CLNDN |
cadd_raw |
cadd_phred |
am_pathogenicity |
mpc |
worst_effect |
gene_effects |
effect_details |
pLI_rank_all |
pLI_rank_min |
LOEUF_rank_all |
LOEUF_rank_min |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
chr11 |
5227002 |
T |
A |
missense |
ENST00000335295.4:HBB:missense:7/147(Glu->Val) |
HBB:missense |
334 |
0.0016 |
0.0127 |
Pathogenic |
HBB-related_disorder|Beta-thalassemia_HBB/LCRB|Fetal_hemoglobin_quantitative_trait_locus_1|Erythrocytosis|_familial|_6|METHEMOGLOBINEMIA|_BETA_TYPE|beta_Thalassemia|Hb_SS_disease|Heinz_body_anemia|Malaria|_susceptibility_to|alpha_Thalassemia|Dominant_beta-thalassemia|Anemia|See_cases|Sickle_cell-hemoglobin_C_disease|Sickle_cell_disease_and_related_diseases|Malaria|_resistance_to|not_provided|Inborn_genetic_diseases|HEMOGLOBIN_S |
1.89 |
18.9 |
0.223 |
0.0374 |
missense |
HBB:missense|ENSG00000298932:non-coding-intron |
ENST00000335295.4:HBB:missense:7/147(Glu->Val)|ENST00000485743.1:HBB:missense:7/111(Glu->Val)|ENST00000647020.1:HBB:missense:7/147(Glu->Val)|ENST00000759072.1:ENSG00000298932:non-coding-intron:None/None[None] |
{‘HBB’: 16189.0} |
1.62e+04 |
{‘HBB’: 19184.0} |
1.92e+04 |
chr12 |
102840493 |
G |
A |
missense |
ENST00000553106.6:PAH:missense:408/452(Arg->Trp) |
PAH:missense |
5030858 |
0.00137 |
0.000861 |
Pathogenic |
PAH-related_disorder|See_cases|Phenylketonuria|Inborn_genetic_diseases|not_provided |
4.22 |
27.9 |
0.918 |
0.245 |
missense |
PAH:missense |
ENST00000307000.7:PAH:missense:403/447(Arg->Trp)|ENST00000553106.6:PAH:missense:408/452(Arg->Trp) |
{‘PAH’: 16912.0} |
1.69e+04 |
{‘PAH’: 15520.0} |
1.55e+04 |
This concludes the Getting Started on the Web section, which demonstrated how to use a saved annotation pipeline in both the Single annotatable and Annotation jobs modes.
The GAIn web Interface section below will describe the web interface in more detail, including custom annotation pipelines, registration, and additional features of the site.