GAIn documentation
Introduction
Annotation is a core step in genomic analysis: sequencing and other genomic assays identify variants, positions, or regions, and annotation adds the biological and clinical context needed to interpret them. GAIn is an infrastructure for running annotation in a consistent, reproducible way—by combining curated resources (genomes, gene models, scores, gene sets, CNV collections, and plugins) with a user-defined annotation pipeline that specifies what to compute and what to emit. This documentation walks through how to use GAIn end to end: getting started on the web and the CLI, discovering and building Genomic Resource Repositories (GRRs), configuring resources, writing and running annotation pipelines, and using the web and Python interfaces.
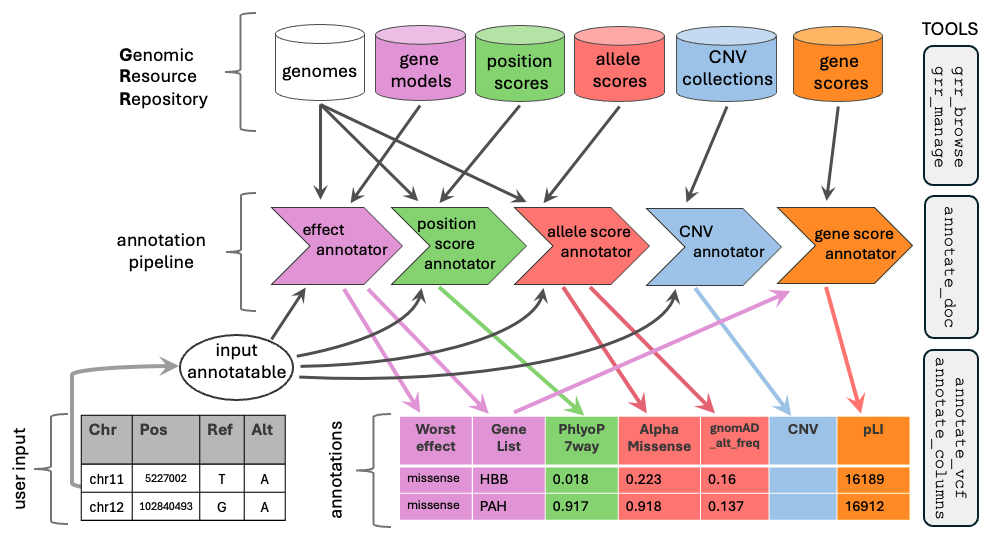
Overview of GAIn. Genomic resources (reference genomes, gene models, scores, CNV collections, gene scores) are organized in a Genomic Resource Repository (GRR). An annotation pipeline chains annotators that draw on these resources to produce an annotated table or VCF. The example highlights dependency flow: a gene score annotator consumes the gene list produced by the effect annotator. Right: core GAIn tools for GRR management (grr_browse, grr_manage), pipeline documentation (annotate_doc), and annotation (annotate_columns, annotate_vcf).